Our Research
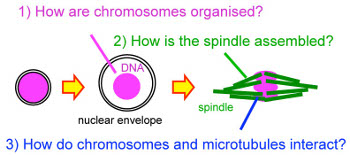
How to build the chromosome segregation machinery
How are genes passed on from cell to cell and from parents to children?
Genes must be passed on accurately from cell to cell and from parents to
children. Failure to do so can be a cause or contributing factor in human
illnesses, such as cancer or reproductive/birth defects. During eukaryotic cell
divisions, cells dramatically change their organisation. DNA carrying genes are
packaged into chromosomes and the spindle made of microtubules is
assembled to segregate the chromosomes. Our lab aims to understand how
chromosomes and the spindle are built and how they interact with each other
at a molecular level using a genetics-led multidisciplinary approach in
Drosophila.
How is the chromosome segregation machinery specialised in oocytes?
One of our recent research focuses has been dissection of molecular pathways
which build the segregation machinery of meiotic chromosomes in oocytes.
Mis-segregation in human oocytes is very frequent and indeed 20% of oocytes
are estimated to mis-segregate chromosomes. This is a major cause of
infertility, miscarriages and birth defects such as Down syndrome. The
chromosome segregation machinery in oocytes shares similarities to that in
mitosis, but also has crucial differences. These differences could be a source of
high error rates in oocytes, but little is known about the molecular pathways
that set up the chromosome segregation machinery specialised to oocytes.
Defining these molecular pathways is crucial to understand error-prone
chromosome segregation in human oocytes. Furthermore, evidence indicates
that apparent oocyte-specific pathways also operate in mitosis, although less
prominently, to ensure the accuracy of chromosome segregation. Therefore
uncovering the molecular basis of these pathways is also important to
understand how somatic cells avoid chromosome instability, a contributing
factor for cancer development. Due to experimental challenges in mammalian
oocytes, we take advantage of Drosophila oocytes as a "discovery platform".
Their similarity to mammalian oocytes and suitability to a genetics-led
mechanistic analysis make them an excellent model system.
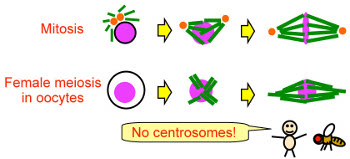
How are microtubules regulated for spindle assembly and function?
Mature oocytes lack centrosomes in many animal species, including humans
and Drosophila. We hypothesise spindle assembly and function in oocytes
should rely more on precise microtubule regulation than in mitosis, even
though oocyte division and mitosis share similar regulatory mechanisms. We
are still far from understanding the full identity and function of microtubule
regulators. Even less is known about how these microtubule regulators
themselves are regulated. By using genetics and 'omics approaches, we have
identified microtubule regulators with functions and regulation pathways in
oocytes that are distinct from mitosis. We are aiming to uncover how oocytes
regulate microtubules to compensate for the absence of centrosomes.
Furthermore, implication of our studies on microtubule regulation is well
beyond cell division, as microtubules are universal features in eukaryotic cells
and involved also in various processes including cell polarity, cell migration and
intracellular transport.
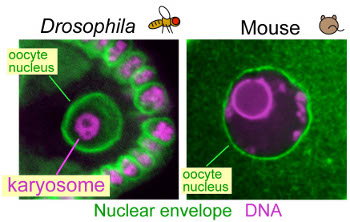
How do chromosomes adopt special organisation in oocytes?
In Drosophila oocytes, meiotic chromosomes cluster together to form a
compact body, called the karyosome within the intact nucleus. A similar
clustering of meiotic chromosomes has also been found in mouse and human
oocytes. In wild-type mouse oocytes, this clustering is known to be correlated
with developmental competency after fertilisation. Based on our studies, we
proposed that the karyosome is crucial for the formation of one unified
acentrosomal spindle. Surprisingly few mechanistic studies on the karyosome
have ever been carried out. We have shown that breaking links between
chromatin and the nuclear envelope is essential for karyosome formation. We
are aiming to uncover the regulatory network which governs the formation of
the karyosome in oocytes.